|
pageCreate()
|
Create a page for a plotgardener layout |
|
pageGuideHide()
|
Remove guides from a plotgardener page |
|
pageGuideHorizontal()
|
Draw a horizontal guideline at a specified y-coordinate on a plotgardener page |
|
pageGuideShow()
|
Reshow guides drawn with pageCreate, pageGuideHorizontal, and pageGuideVertical |
|
pageGuideVertical()
|
Draw a vertical guideline at a specified x-coordinate on a plotgardener page |
|
pageLayoutCol()
|
Generate column positions for a number of plot elements with a specified width and space between them |
|
pageLayoutRow()
|
Generate row positions for a number of plot elements with a specified height and space between them |
|
pagePlotPlace()
|
Place a plot that has been previously created but not drawn |
|
pagePlotRemove()
|
Remove plotgardener plots and annotations |
 |
plotGenes()
|
Plot a gene track for a specified genomic region |
 |
plotGenomeLabel()
|
Plot genomic coordinates along the x or y-axis of a plotgardener plot |
 |
plotHicRectangle()
|
Plot a triangular Hi-C interaction matrix in a rectangular format |
 |
plotHicSquare()
|
Plot a Hi-C interaction matrix in a square format |
 |
plotHicTriangle()
|
Plot a Hi-C interaction matrix in a triangular format |
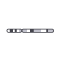 |
plotIdeogram()
|
Plot a chromosome ideogram with or without cytobands |
 |
plotManhattan()
|
Plot a Manhattan plot |
 |
plotPairs()
|
Plot paired-end genomic range elements |
 |
plotPairsArches()
|
Plot paired-end genomic range data in an arch style |
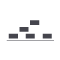 |
plotRanges()
|
Plot genomic range elements in a pileup or collapsed format |
 |
plotSignal()
|
Plot any kind of signal track data for a single chromosome |
|
plotMultiSignal()
|
Plot multiple signal tracks in line with each other |
 |
plotTranscripts()
|
Plot gene transcripts in a pileup style for a single chromosome |